The database "fanta.bio" that contains information on genomic regions and cis-elements involved in gene transcription regulation is now available.
- Others
- Funding
- DICP Tool Prototype Trial
On February 26, 2024, "fanta.bio," a database of cis-regulatory elements (CREs), which are genomic regions involved in the transcriptional regulation of genes, was officially released.
Cis-elements are regions in the genome that are involved in transcriptional regulation, and typical examples include enhancers and promoters. The fanta.bio contains information on
- the location of cis-elements identified in the human and mouse genomes,
- the activity of cis-elements in different cell lines, cell types and tissues,
- genomic variations within cis-element regions.
Using fanta.bio, you can search for cis-elements in any genomic region or gene, and obtain information on the activity of the cis-element in cells (whether the cis-element exerts regulatory activity in that celll, etc.), transcription factors that bind to the cis-element region, genomic polymorphisms in the region, etc. As well as the search results can be browsed through the interface provided on the website, the genomic location can be confirmed using the UCSC Genome Browser. In addition, data files created by fanta.bio can be downloaded. By using these data files, it is possible, for example, to integrate and analyze data such as genomic mutation information related to a disease, and to investigate the relationship between cis-elements and the disease.
In the construction of fanta.bio, information on transcription factor binding regions was integrated from the epigenomics database ChIP-Atlas, and genomic polymorphism information on mice was integrated from MoG+, which collects strain-specific genomic variation for mouse genome polymorphism information. For human genome polymorphism information, TogoVar, developed and operated by DBCLS, is being used.
In the future, it will be possible to collectively obtain information on transcriptional regulatory regions by enhancing activity information in more diverse cell types using single-cell gene expression data and various related genome function information. In addition, the linkage between fanta.bio and ChIP-Atlas will be further strengthened, and the integrated infrastructure related to transcriptional regulation, INTRARED, will be enhanced.
<Number of data included in fanta.bio (ver. 1.0.0)>
- Human (447,315 CRE regions, 1,912 cis-element activities)
- Mouse (201,420 CRE regions, 1,052 cis-element activities)
The fanta.bio, together with ChIP-Atlas, is a project of the JST Database Integration Coordination Program (DICP) "Construction of an integrative data platform for transcriptional regulations (Principal Investigator: Takeya Kasukawa, Team Leader, RIKEN)"
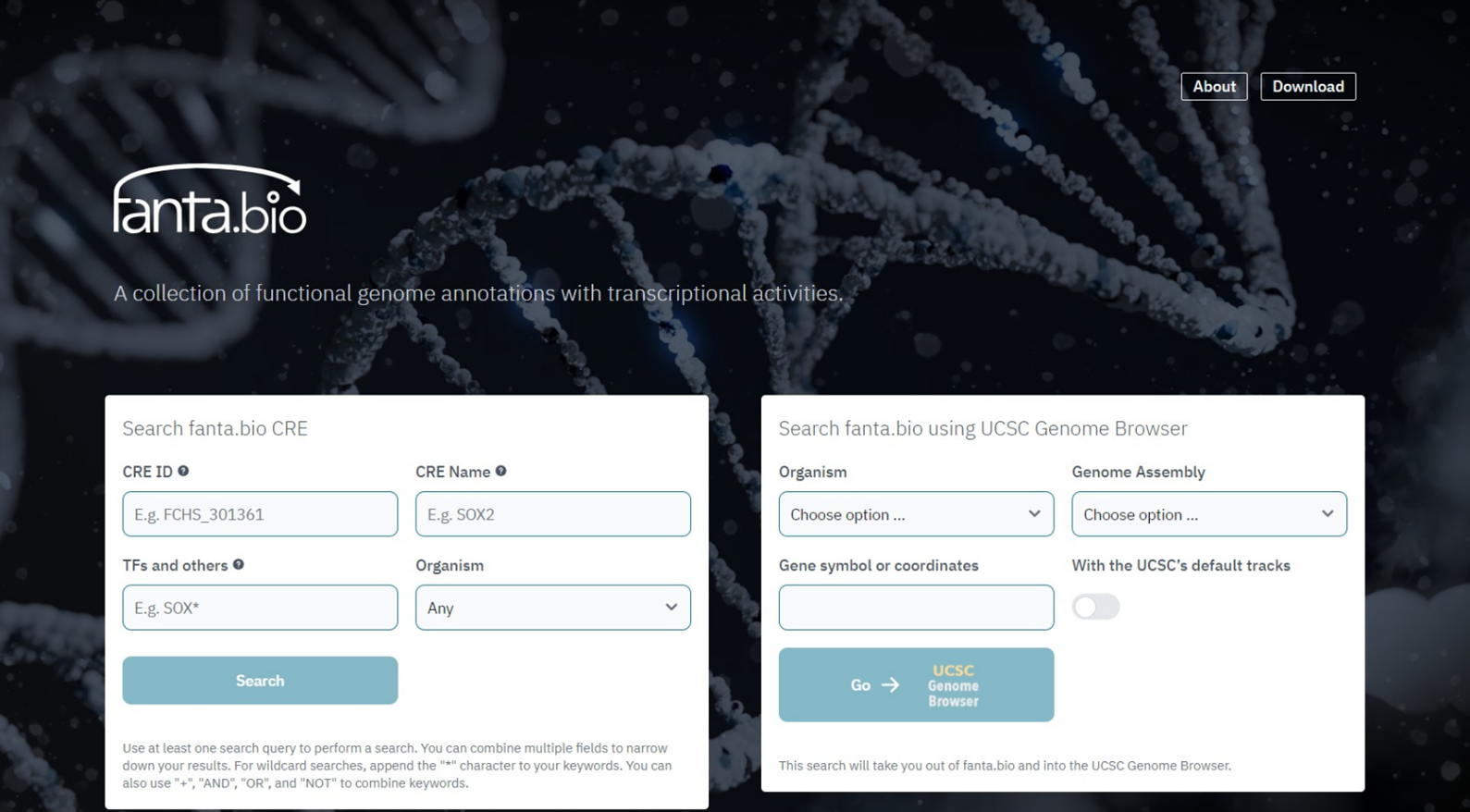
Search page of fanta.bio
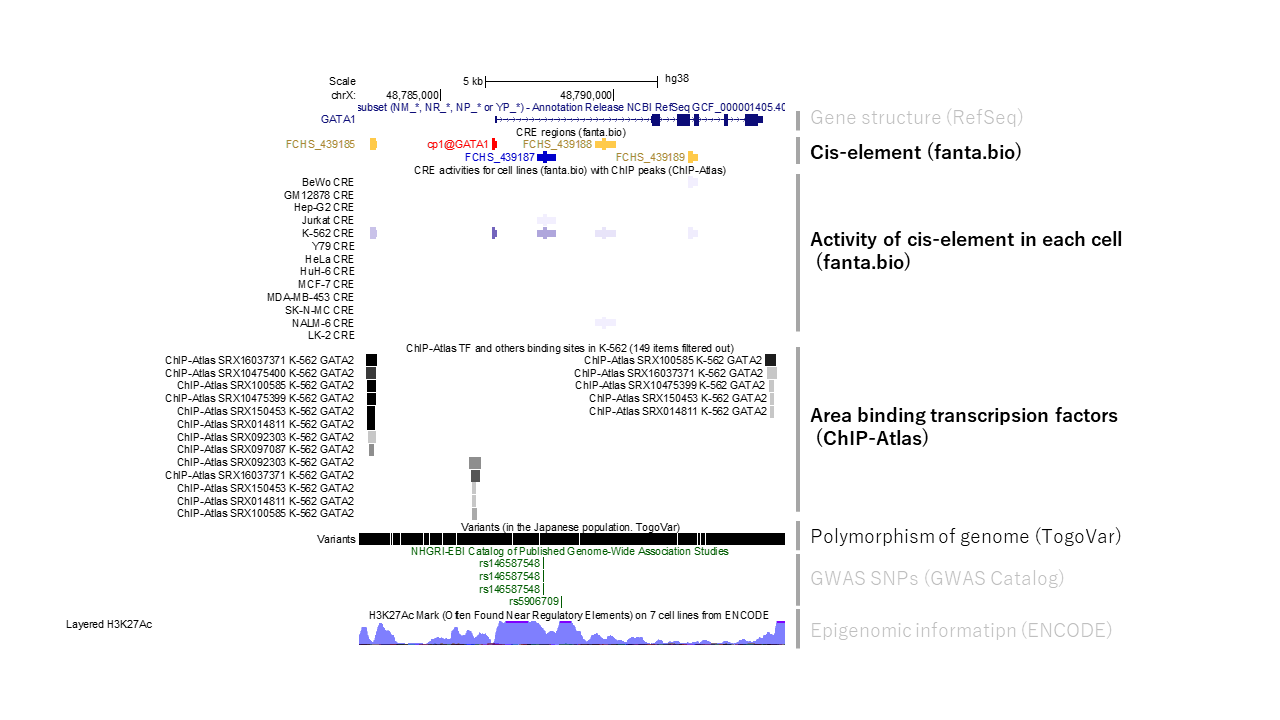
Human cis-element (promoter and enhancer region of GATA1 gene)

Mouse cis-element (promoter region of Gata1 gene)
Related Links
- Cis-element database fanta.bio
- Integrative epigenome database ChIP-Atlas
- INtegrated TRAnscriptional REgulation Data platform INTRARED
- Construction of an integrative data platform for transcriptional regulations - Adopted Proposals | NBDC website
Outline of the proposal, etc.