A paper on Microbiome Datahub, an integrated database that collects and curates genome sequences derived from metagenomic analyses of environmental microorganisms, has been published
- Others
- Funding
- Database Integration Coordination Program
Dr. Hiroshi Mori, Associate Professor of National Institute of Genetics (NIG), Research Organization of Information and Systems, and his colleagues published a paper in the advance online edition of the scientific journal "Microbiome" describing the development of "Microbiome Datahub", an integrated database created by comprehensively collecting metagenomic genome sequences (MAG: Metagenome-Assembled Genomes) derived from analysis of microorganisms in the environment from public nucleotide sequence repositories, and adding various information such as environment, phylogeny, and gene function on Mar 16, 2026. On Mar 30, NIG, the National Institute for Basic Biology (NIBB) of the National Institutes of Natural Sciences, Institute of Science Tokyo, and Kyoto University issued a joint press release highlighting the significance of Microbiome Datahub. The press release states that Microbiome Datahub will contribute as a foundation for data-driven microbiology and the discovery of novel beneficial proteins.
With the development of metagenomic analysis technology that directly decodes DNA sequences from the microbiome (*1) in the environment, the amount of metagenomic genomic information (MAG) (*2) from microorganisms that are difficult to culture has increased explosively. Although this data is registered in public DNA sequence repositories (INSD) (*3), there are challenges such as variations in quality, unorganized environmental metadata (habitat), lack of genetic information, and inconsistent classification systems, making it difficult to search the data or perform cross-sectional comparative analysis. In addition, existing secondary databases of MAGs (MGnify, IMG/M, SPIRE, etc.) reconstruct MAGs independently, which can result in sequences that differ from those reported in original research papers, making it difficult to correctly refer to and evaluate the research results of original papers. To solve these problems, Associate Professor Mori and his colleagues developed "Microbiome Datahub", an integrated database of MAGs that adopts an approach that unifies and enhances only the metadata and annotations while keeping the MAG sequences in public repositories intact.
Microbiome Datahub collected 214,427 MAGs from INSD and organized environmental metadata using a proprietary ontology (MEO: Metagenome and Microbes Environmental Ontology) that systematically describes the habitat of microorganisms. Approximately 170,000 of the included MAGs met high quality standards of Completeness >60% and Contamination <10%. In addition, using the prokaryotic phenotype prediction tool "Bac2Feature" which uses phylogenetic names and 16S rRNA gene sequences, 27 phenotypes, including growth rate, optimal temperature, and optimal pH, were predicted for all MAGs from the phylogenetic name and added as information to the MAGs. Analysis using the ultra-fast sequence similarity search tool "PZLAST" revealed that approximately 19% of the protein sequences encoded in the MAGs included in Microbiome Datahub are highly novel protein sequences that do not have homology to existing orthologs (*4) databases (MBGD).
Microbiome Datahub offers high-speed web-based searching, API access, and bulk downloads, and is expected to be widely used in applications ranging from basic microbiology research to applied research such as protein structure prediction and useful enzyme discovery. Microbiome Datahub is planned for continuous updates and expansion as a database that will contain, organize, and publish the rapidly growing public MAG data.
For more details, please refer to the paper and the press release.
< Number of data points included in Microbiome Datahub (ver. 1.0)>
- MAG: 218,248 MAGs
- Metagenomic BioProject: 102,174 projects
- MAG-derived protein sequences: 454,799,346 proteins
Microbiome Datahub is developed and provided as part of JST Database Integration Coordination Program (DICP), "Development of an integrated microbiome data hub for microbiome research" (Principal Investigator: Associate Professor MORI Hiroshi, National Institute of Genetics). It is achieved through collaboration with "MBGD" (Microbial Genome Database for Comparative Analysis), an ortholog database of microorganisms, and "Bac2Feature", a tool for estimating microbial phenotypes, both of which are being developed by the research group of this project.
Terminology
*1 Microbiome: It refers to the community of microorganisms that inhabit environments such as soil, water, the surface of the body, and the intestines. Microbiome is composed of a wide variety of microorganisms and influences the surrounding environment while maintaining a delicate balance. Microbiome of cultivated land soil is thought to interact with crops and influence crop growth and yield, and analysis of the soil microbiome is expected to contribute to improving production efficiency. Furthermore, the gut microbiome is thought to influence health and disease, and its analysis is expected to contribute to healthcare by elucidating disease onset mechanisms and developing preventive and therapeutic methods.
*2 MAG (Metagenome Assembled Genome): A hypothetical genome sequence obtained by extracting DNA from a sample as a mixture without culturing a microbial community, comprehensively sequencing the base sequence to obtain a metagenomic, assembling the resulting metagenomic sequence, and then clustering (binning) the sequences based on information such as the sequence's continuous base composition and relative abundance from the resulting contig sequences.
*3 INSD (International Nucleotide Sequence Database): A public DNA sequence repository that accumulates and publishes DNA sequence data of various organisms. Currently, it is operated by three institutions: the National Institute of Genetics (DDBJ), ENA at EMBL-EBI in Europe, and NCBI in the United States.
*4 Orthologs: They refer to the correspondence between genes that have evolved from a common ancestral gene, or to groups of genes that have such correspondences. Orthologs generally have similar functions and are important clues for comparative studies of gene functions between species and for estimating the function of newly discovered genes.
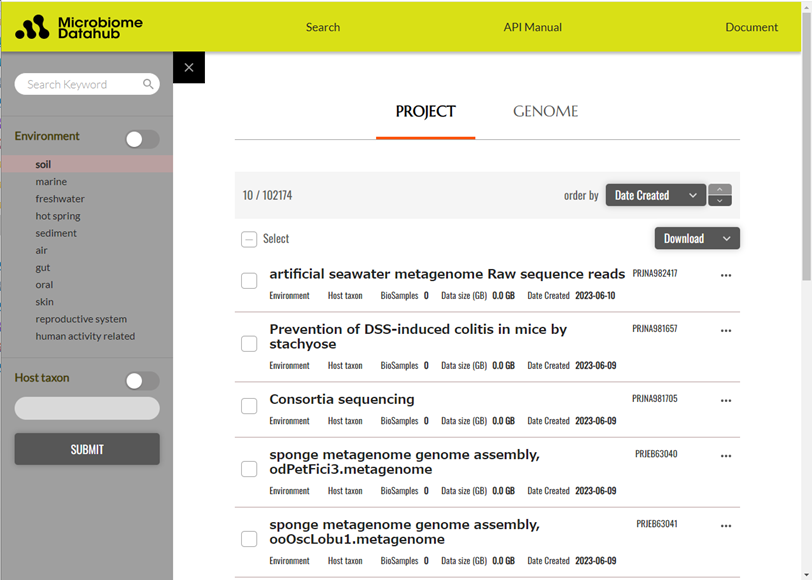
Figure 1. Search page for project information in Microbiome Datahub
In addition to searching for projects using free keywords, you can also refine your search by specifying the environmental information from which the analyzed samples were obtained (soil, marine, freshwater, hot spring, sediment, air, gut, oral, skin, reproductive system, etc.), or, in the case of samples derived from animal intestines, information such as the host taxon's lineage.
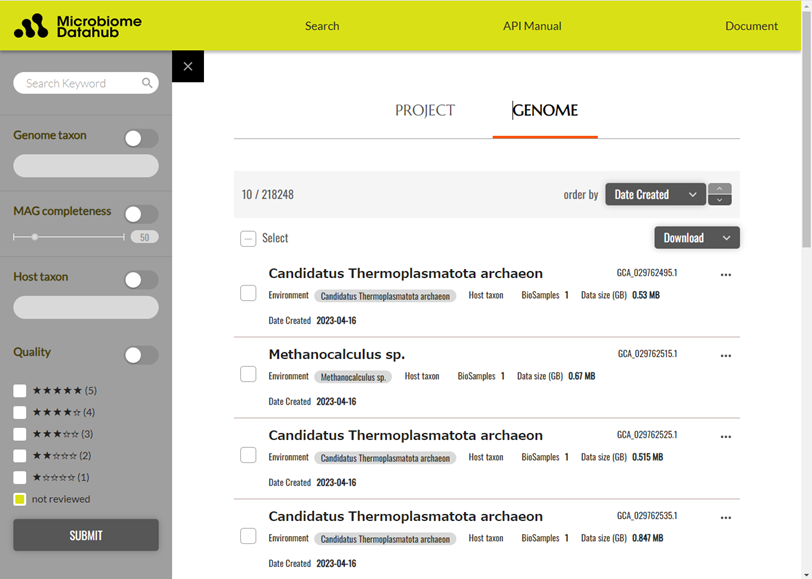
Figure 2. Search screen for genome information in Microbiome Datahub
In addition to searching genome information using free keywords, you can refine your search based on information such as the estimated microbial taxon of the MAG, the MAG's quality, the host taxon (if the MAG originates from animal intestines, etc.), and the MAG's quality.

Figure 3. Statistics of DFAST and DFAST_QC for MAG, phenotypic estimation results of Bac2Feature, and an example of MBGD Ortholog composition display (Bifidobacterium catenulatum GCA_022728695.1)
You can view metadata of the MAG, statistics of the MAG, Quality evaluation of the MAG, phenotype estimation by Bac2Feature, assigned MBGD Ortholog list and KEGG Ortholog list.
Related Links
- "Microbiome Datahub: an open-access platform integrating environmental metadata, taxonomy, and functional annotation for comprehensive metagenome-assembled genome datasets." | Microbiome
- Press release "Development of "Microbiome Datahub," an integrated database compiling and organizing metagenome-derived genomes"|National Institute of Genetics (NIG)
- Microbiome Datahub
- Development of an integrated microbiome data hub for microbiome research | NBDC
Project summaries and reports are posted.