Plant co-expression database "ATTED-II" update: plant species has been expanded to 19 species - 20 lineages, and a new inter-species comparative analysis tool has been implemented
- Others
- Funding
- Database Integration Coordination Program
On Feb 18, 2026, Professor Takeshi Obayashi and his colleagues at the Graduate School of Information Sciences at Tohoku University updated the plant gene co-expression database "ATTED-II" to v13, expanding the number of plant species containing co-expression information to 19 species - 20 lineages, and implementing a new analytical tool for comparing co-expression relationships across species.
In recent years, genome analysis has been conducted in a variety of plant species, including practical plants such as grains and vegetables. in contrast to "model plants" such as Arabidopsis that have been well studied up until now, many genes whose functions are unknown have been discovered from them. Gene groups whose expression is correlated under specific conditions or environments often play a role in a series of related physiological functions, and co-expression information is thought to be useful in understanding the functions of these genes and the regulatory relationships of their expression.
ATTED-II was first released in 2004 as a database containing plant co-expression information. In the formaer update to v12 in Sep 2024, the number of plant species was expanded to 11, and the following features were added: "PC View," which allows users to investigate the conditions and environments in which two genes of interest are co-expressed through principal component (PC) analysis of co-expressed genes, and "CoexViewer," which visualizes the correlation diagram of the expression patterns of two genes of interest.
This update to ATTED-II v13 expands the range of plant species to 19 species - 20 lineages. And also it implements new interspecies comparison analysis tools, enabling comparison of co-expressed genes across species and further insight into their physiological functions. The "PC View," which displays the sample environment based on principal component analysis of a large number of samples, has been enhanced to display not only sample and gene lists related to each principal component axis, but also provides summaries, making it easier to examine differences across multiple principal component axes. Specifically, KEGG enrichment tests are now provided for gene lists, and the results and variable samples are summarized using LLM at three levels of granularity: labels, short summaries, and long summaries. Labels and short summaries are also displayed in the "Coex Viewer," which displays the co-expression of gene pairs, providing a new feature to aid in understanding co-expression conditions. Furthermore, the interspecies comparison function for co-expression information has been enhanced, allowing users to compare and analyze similarities and differences in co-expressed genes between distantly related species by displaying co-expression maps by subcellular location, such as chloroplasts and nuclei.
ATTED-II is developed as a part of JST Database Integration Coordination Program (DICP), "Platform of gene network information for non-model plants" (Principal Investigator: OBAYASHI Takeshi, Professor, Graduate School of Information Sciences, Tohoku University).
Plants included in ATTED-II v13 (Total 19 species - 20 lineages)
<Dicotyledon>
- Arabidopsis - standard lineage(Arabidopsis thaliana; ath-r)
- Arabidopsis - non-standard lineage(Arabidopsis thaliana; ath-e)
- Tomato(Solanum lycopersicum; sly)
- Medicago(Medicago truncatula; mtr)
- Soybean(Glycine max; gma)
- Poplar(Populus trichocarpa; ppo)
- Grape(Vitis vinifera; vvi)
- Field mustard(Brassica napus; bna)**
- Upland cotton(Gossypium hirsutum; ghi)**
- Potato(Solanum tuberosum; sot)**
- Tobacco(Nicotiana tabacum; nta)**
- Orange(Citrus sinensis; cit)**
<Monocots>
- Japonica rice(Oryza sativa subsp. japonica; osa-r)
- Indica rice(Oryza sativa subsp. indica; osa-e)**
- Maize(Zea mays; zma)
- Wheat(Triticum aestivum; tae)*
- Barley(Hordeum vulgare; hvu)*
- Sorghum(Sorghum bicolor; sbi)**
- Brachypodium(Brachypodium distachyon; bdi)**
<Green algae>
- Chlamydomonas(Chlamydomonas reinhardtii; cre)**
* indicates plant species for which co-expression information was newly added in v12 (Sep 2024).
** indicates plant species and strains for which co-expression information was newly added in v13 (Feb 2026).

Fig 1 Top page of ATTED-II v13
The ATTED-II v13.0 top page. It features links to search functions (Search) such as GeneTable, EdgeAnnotation, and CoExSearch, links to analysis functions (Draw) such as NetworkDrawer, CoexMap, and Hcluster, links to main sample pages for each of the 19 plant species and 20 lineages included (Target species with page examples), and links to the download page (Bulk download).
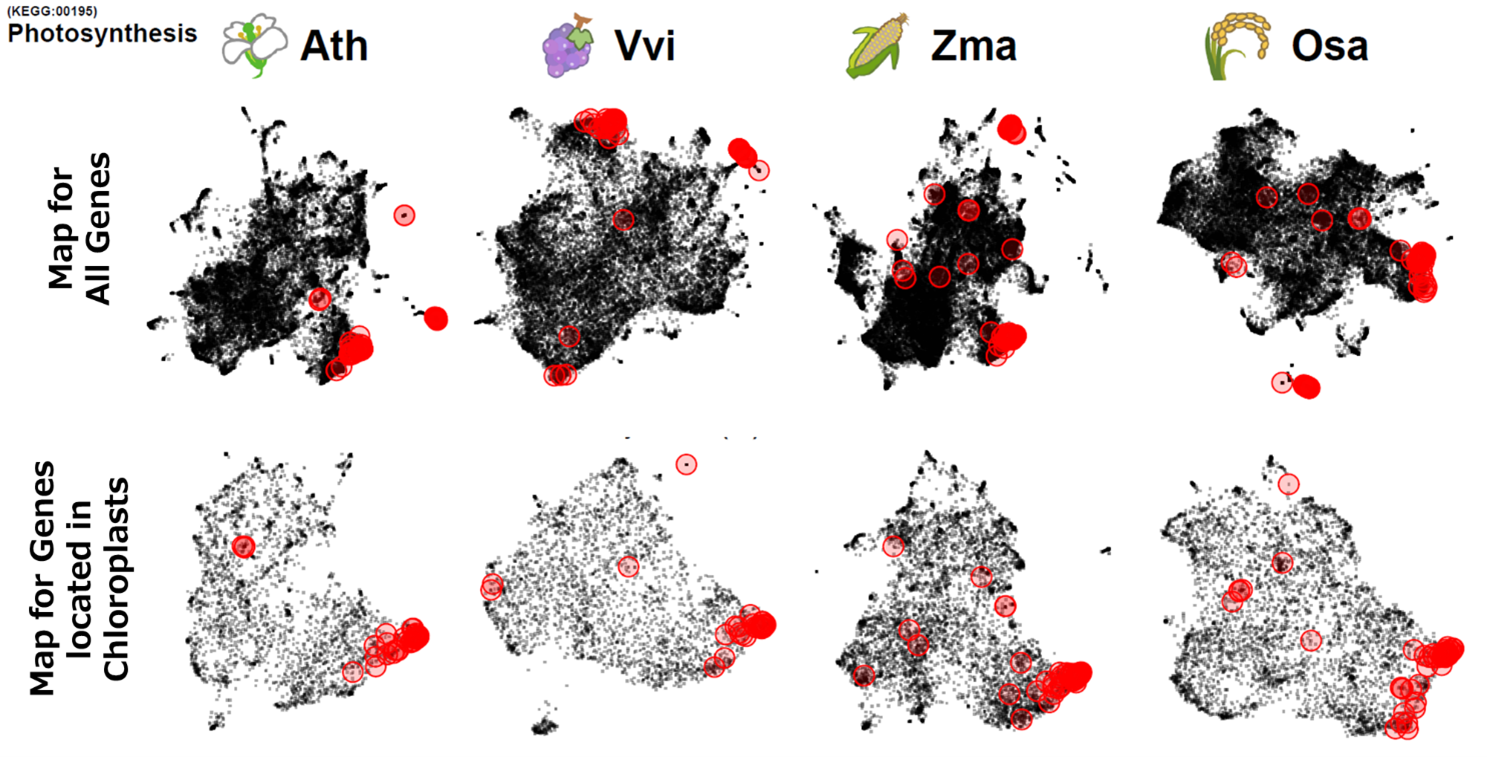
(A) Photosynthesis-related genes in Chloroplasts
(B) Ribosome biogenesis-related genes in Nucleus
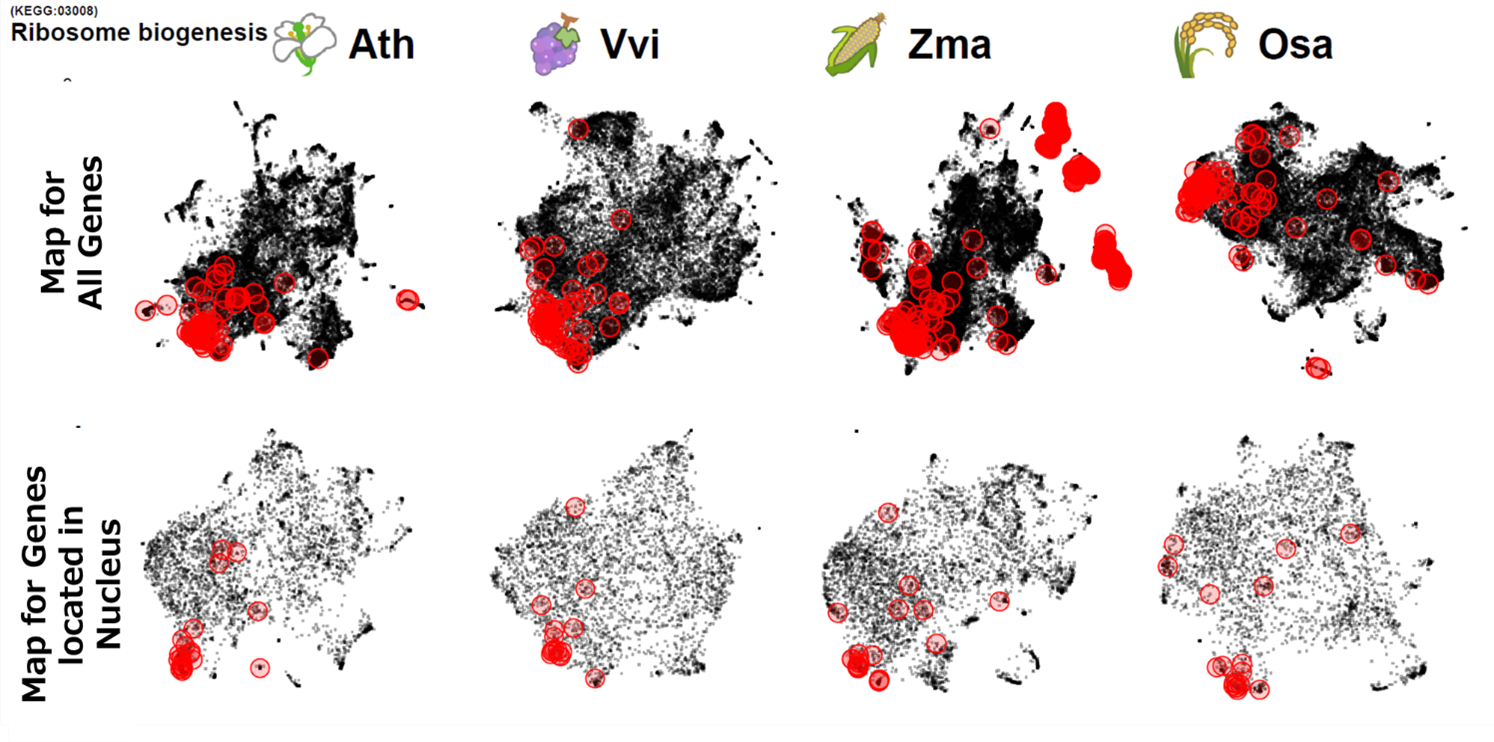
Fig 2 Analysis tool for comparing co-expression relationships across species
(A) Comparison of genes co-expressed in chloroplasts between plant species. From left: Arabidopsis (Ath), grapevine (Vvi), maize (Zma), rice (Osa). Red circles show photosynthesis-related genes (KEGG:00195). The top row shows a map of co-expressed genes across all genes, while the bottom row shows a map of co-expression genes for proteins localized in chloroplasts. Looking at all genes, no consistency was observed in the localization of photosynthesis-related genes (KEGG:00195), but a consistent location was observed between species when focusing on chloroplast localization.
(B) Comparison of genes co-expressed in nuclei between plants. From left: Arabidopsis (Ath), grapevine (Vvi), maize (Zma), rice (Osa). Red circles show ribosome biogenesis-related genes (KEGG:03008). The top row shows a map of co-expressed genes across all genes, while the bottom row shows a map of co-expression genes for proteins localized in the nucleus. Looking at all genes, no consistency was observed in the localization of ribosome biogenesis-related genes (KEGG:03008), but a consistent location was observed between species when focusing on nuclear localization.
関連リンク
- ATTED-II
- Funded projects: "Platform of gene network information for non-model plants" | NBDC
Outline of the project, etc.